My name is Priscilla San Juan. I am a data science specialist at the
Natural History Museum of Los Angeles County. I earned my PhD
in Ecology & Evolution from Stanford University and have over a decade of experience designing
experiments, generating end-to-end analytical pipelines, and communicating
findings to diverse audiences.
My research background has given me the expertise in the kinds of problems
that matter in industry: understanding patterns through time,
segmenting data into meaningful groups, building reproducible models
that scale, and translating complex outputs into clear recommendations.
As a native Angeleno, I share a love for tv and film. I am excited to
apply my skills to understand audience behavior, content
performance, and viewer engagement as a way to contribute to a creative industry I admire.
Below, I would like to showcase some of my work with data.
Projects
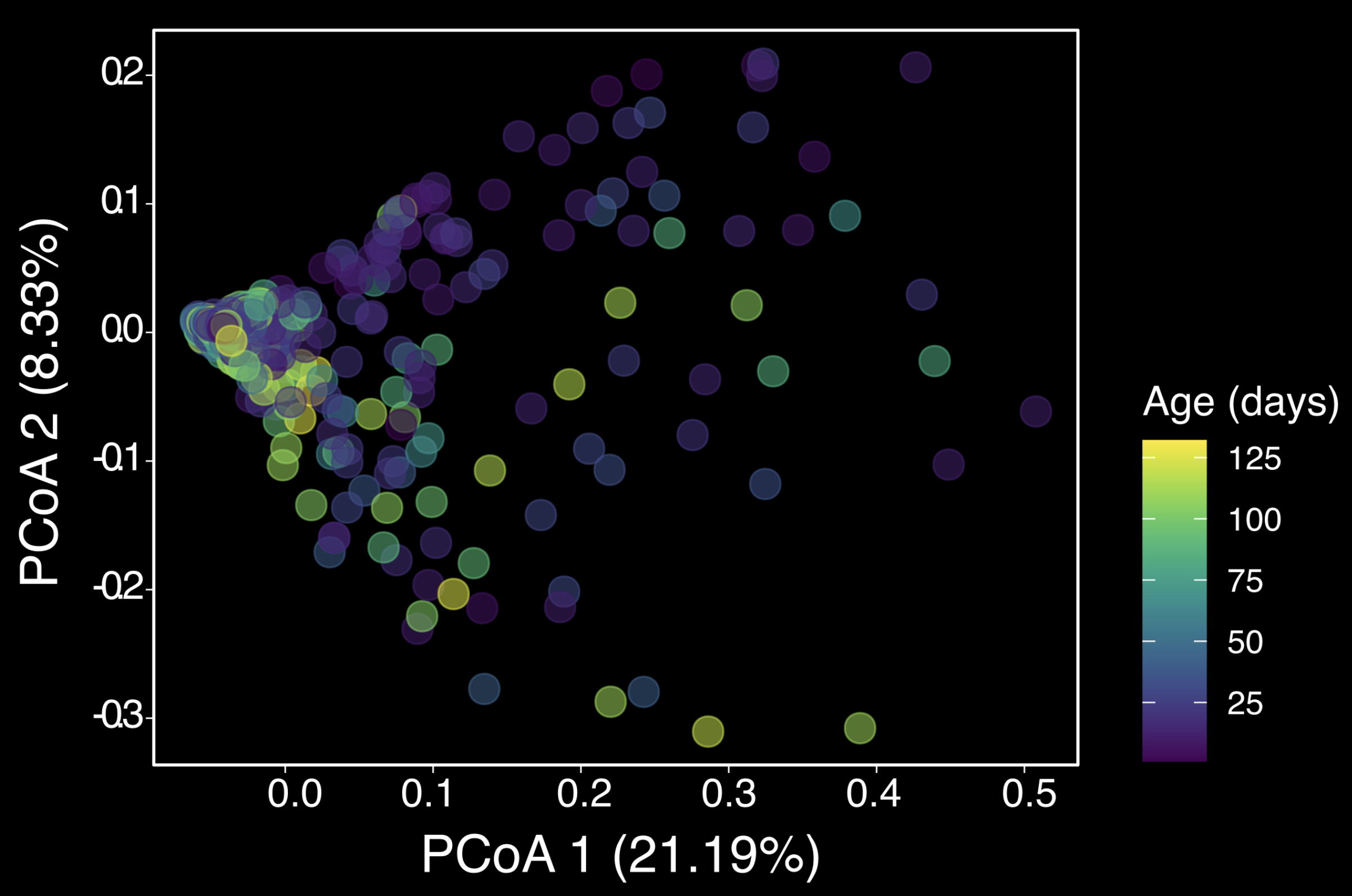
Time-series analysis of complex communities
In this study, I demonstrate how longitudinal data analysis can uncover structure and
dynamic trends in complex biological systems. By collecting over 800 microbiome samples
and integrating them with environmental (soil) and health metadata, the project applied
advanced statistical modeling, diversity metrics, and multivariate ordination to track
microbiome change over time in developing kiwi. The result was a high-resolution model of
microbiome assembly that identifies key environmental drivers and temporal stabilization
patterns.
Bioinformatics script, R analysis code, and processed data
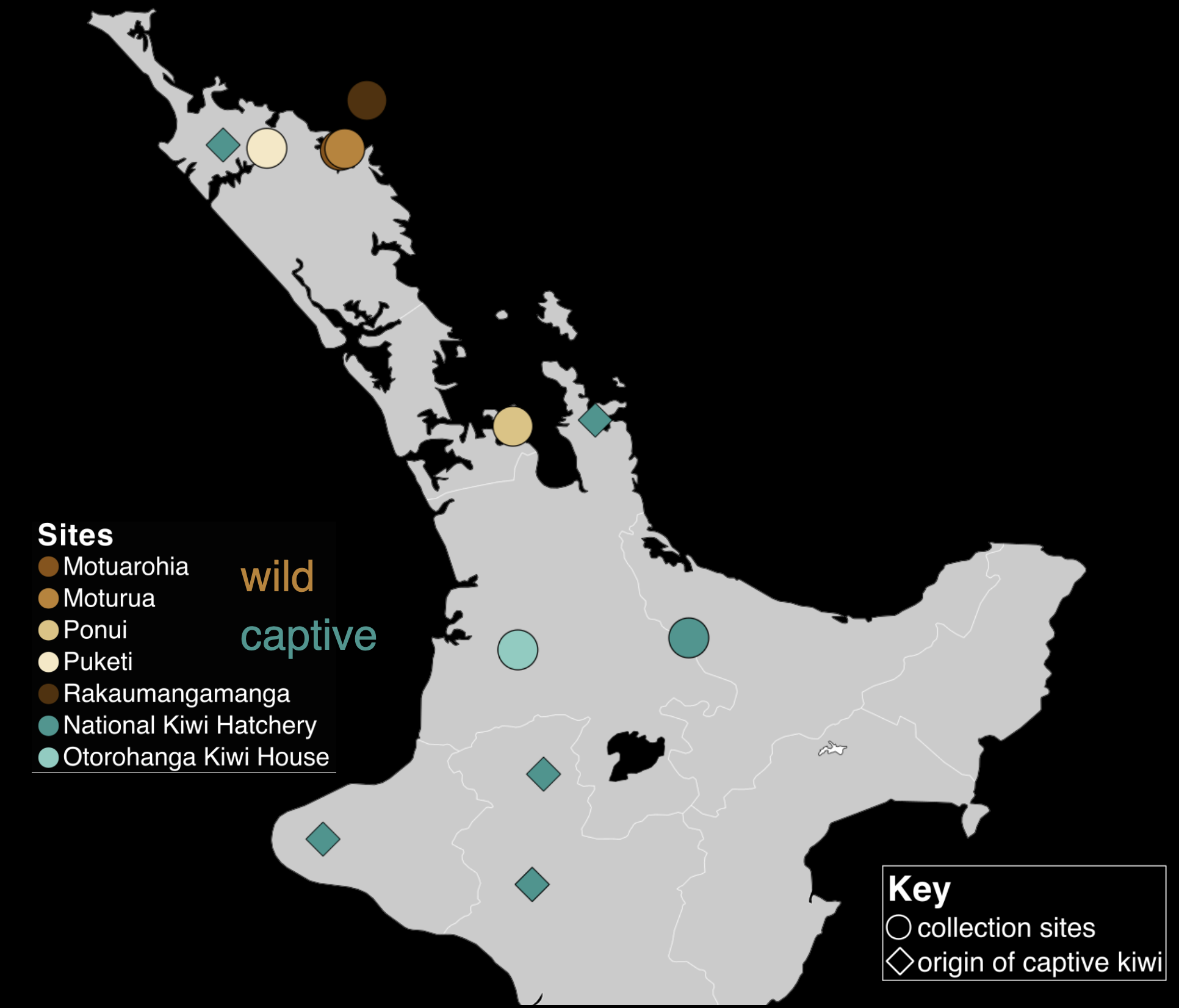
Determining the effect of captive-rearing on Brown Kiwi
This study combined next-generation sequencing data with advanced statistical analysis to
assess how captivity alters the gut microbiome of an endangered animal, the Brown Kiwi.
By integrating large-scale biological datasets, spatial analysis, and compositional
clustering, the project demonstrates how robust analysis can extract valuable insights
from complex, noisy data.
Processed data
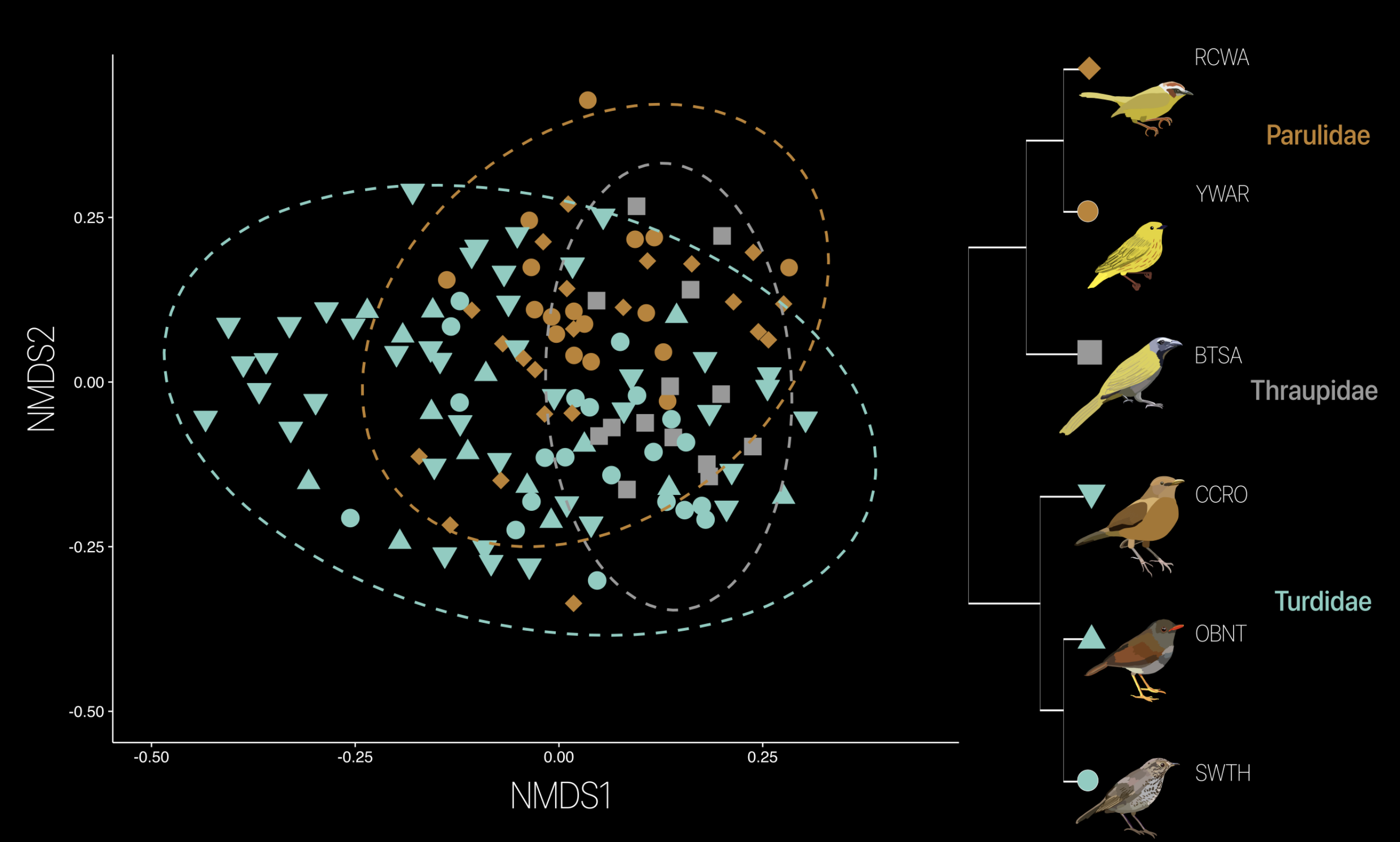
Analyzing what factors shape bird gut microbiomes
Here, I showcase how environmental analytics can uncover complex relationships between
biological systems (i.e. birds) and human-modified environments. By integrating 280 microbiome
datasets from six bird species across a land-use gradient in Costa Rica, the research applied
multivariate modeling, interaction testing, and compositional data analysis to detect subtle
but significant effects of habitat transformation on gut microbial diversity. The result is a
reproducible framework for quantifying how environmental change impacts host-associated microbiomes.
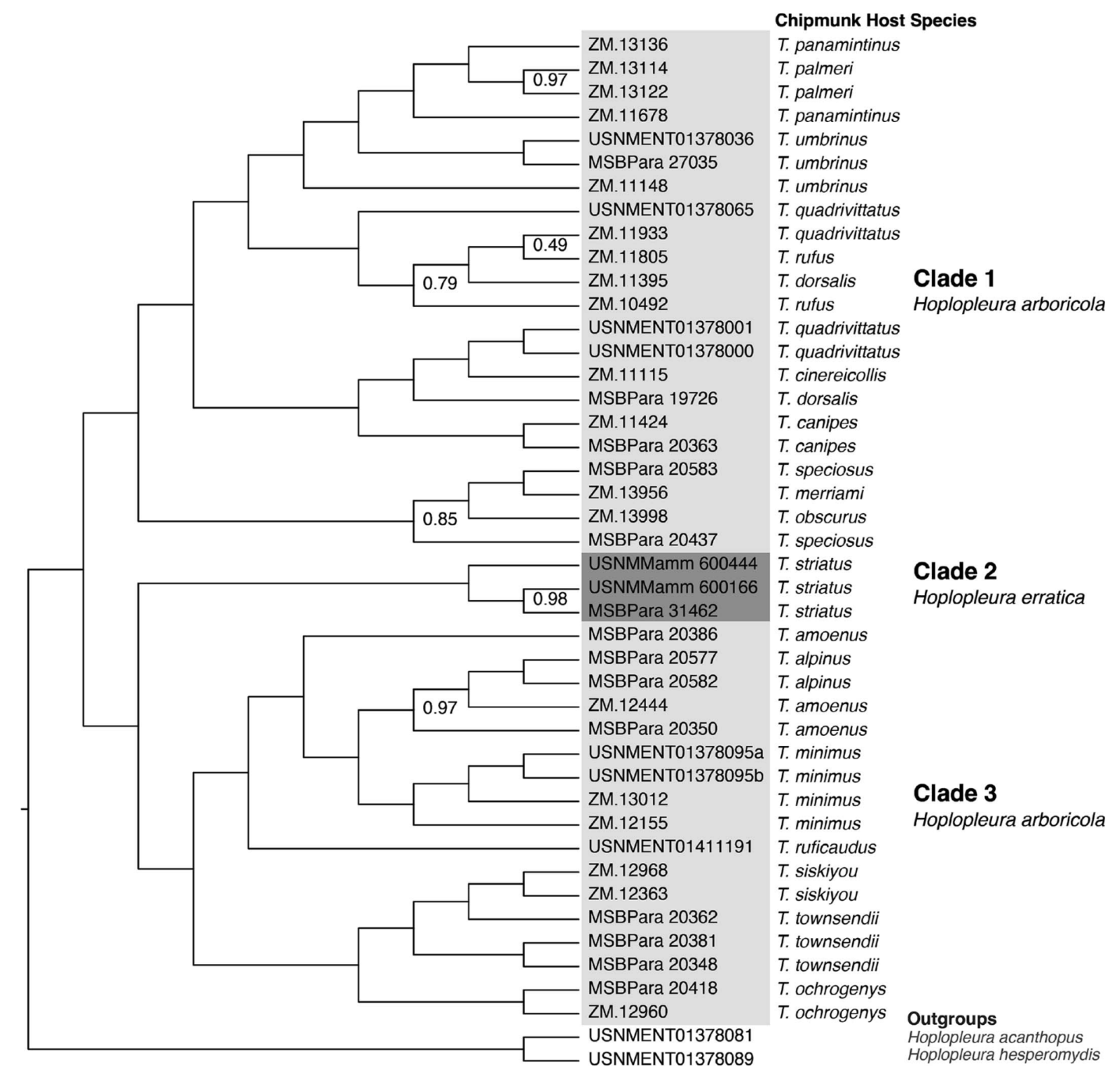
Resolving taxonomy with data
This study exemplifies how data analysis can transform complex datasets into
actionable insights, in this case taxonomic revisions. By integrating over a thousand
genomic loci with morphological and host-association
data, the research resolved long-standing classification challenges in parasitic insects.
Skills
Languages & Tools:
R, Python, SQL, Excel, Google Sheets, Bash, Git, Jupyter
Data Visualization:
ggplot2, Tableau
Statistical Analysis:
Supervised and unsupervised learning, regression modeling, time-series analysis, spatial analysis
Data Management:
Data validation, relational database querying, large-scale data integration, ETL pipelines
Other:
QGIS, Linux/Unix environments, high performance computing, Asana, Sharepoint, Illustator




